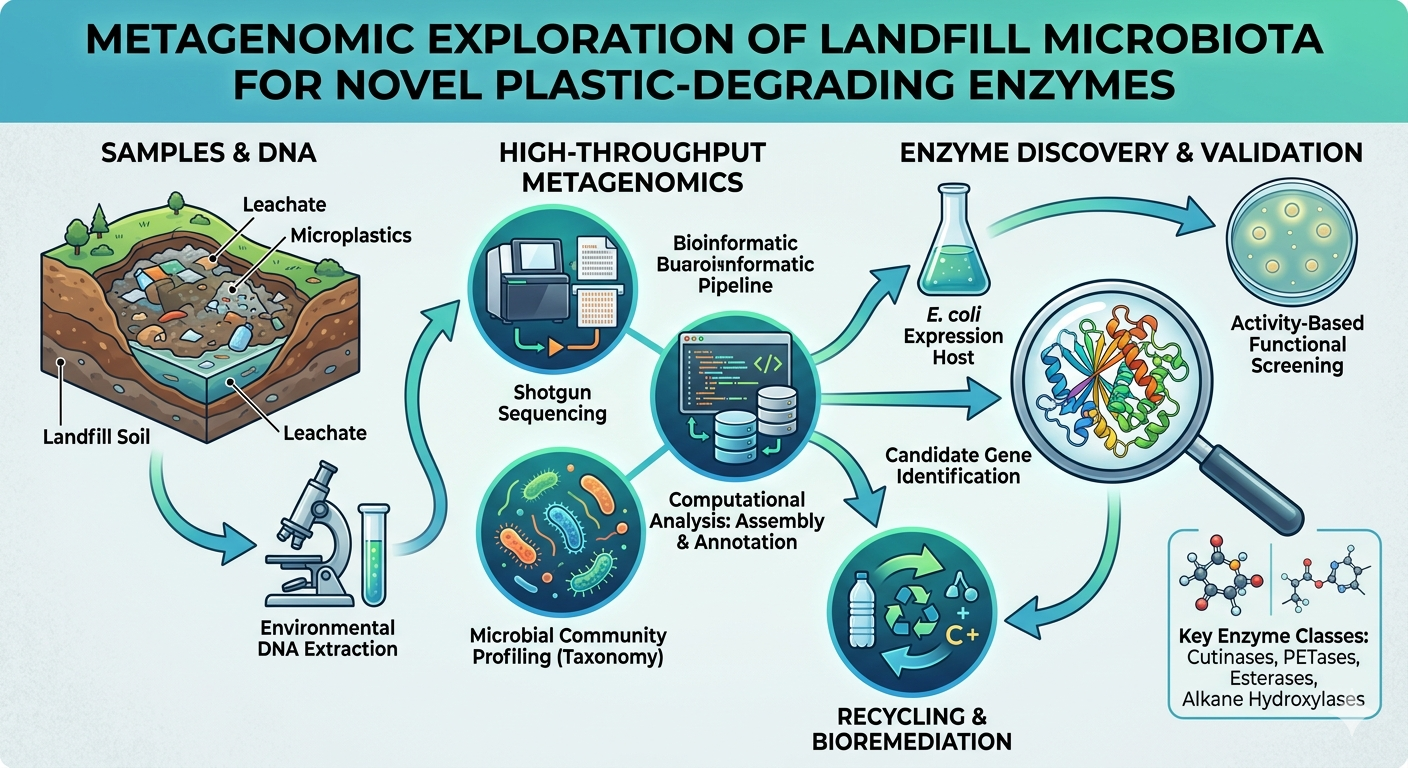
Metagenomic Exploration of Landfill Microbiota for Novel Plastic-Degrading Enzymes
DOI:
https://doi.org/10.64062/JPGMB.Vol2.Issue2.8Keywords:
- Plastic pollution; Metagenomics; Landfill microbiota; Plastic-degrading enzymes; PETase; Cutinase; Biodegradation; Microbial consortia; Bioinformatics; Sustainable waste management.
Abstract
Plastic waste has become one of the most pressing environmental challenges worldwide, primarily because synthetic polymers do not easily break down under natural conditions. As a result, large quantities of plastic accumulate in landfills, where they persist for long periods. Interestingly, these landfill environments are not just waste storage sites but also complex ecosystems that host diverse microbial communities. Over time, some of these microorganisms have adapted to survive by interacting with and even utilizing resistant plastic materials. Recent advancements in metagenomics have made it possible to study these microbial communities without the need for traditional culturing techniques. This has opened new avenues for discovering previously unknown plastic-degrading enzymes. In this review, we explore how landfill microbiota act as valuable sources of biodegradation potential and discuss various metagenomic approaches used to identify such enzymes. Special attention is given to key enzyme groups, including PETases, cutinases, lipases, and esterases, which play crucial roles in breaking down synthetic polymers. This review also outlines these limitations and discusses future directions for developing sustainable and effective plastic waste management strategies.
References